
The R.1 isolates clustered on their own distinctly from B.1.1.7 (Alpha), B.1.351 (Beta), B.1.617.2 (Delta), P.1 (Gamma), and B.1.1.529 (Omicron) variants ( Figure 1). 
We performed phylogenetic analysis to confirm clustering with the R.1 lineage. We determined the viral whole-genome sequences and deposited them into GISAID (accession nos. The 2 R.1 isolates were purified from nasopharyngeal samples from SARS-CoV-2–specific qRT-PCR–positive persons by using Vero clone E6 cells as described previously ( 10). We isolated 2 SARS-CoV-2 isolates belonging to the R.1 lineage (R.1 645, R.1 646) from 2 patients from Hamilton, Ontario, Canada, in March 2021, corresponding to the third wave of the COVID-19 pandemic in Canada. Phylogenetic confirmation that SARS-CoV-2 isolates belong to R.1 lineage in study of sensitivity to neutralizing antibodies and resistance to type I interferons in SARS-CoV-2 R.1 lineage variants, Canada. 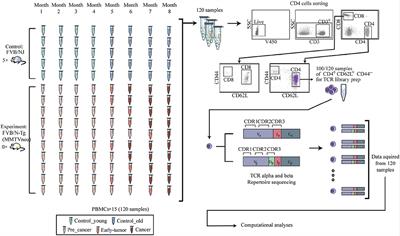
Therefore, we evaluated the sensitivity of SARS-CoV-2 R.1 lineage isolates from patients in Canada to neutralizing antibodies and type I interferons.įigure 1. Taken together, characterizing IFN-resistant SARS-CoV-2 variants is critical, given their potential to enhance transmission kinetics and result in viral evolution. Furthermore, SARS-CoV-2–infected persons with genetic defects in IFN signaling are at higher risk for severe COVID-19 ( 9). A recent study compared multiple type I IFNs against diverse SARS-CoV-2 VoC, demonstrating increased IFN resistance ( 8). SARS-CoV-2 proteins are involved in IFN evasion either by directly suppressing production or by acting downstream of the host response machinery ( 6, 7). The type I interferon (IFN) response constitutes the first line of defense against many viruses ( 5, 6), triggering activation of several IFN-stimulated genes (ISGs) and establishing an antiviral state ( 5). However, data on the immune-evasive properties of this lineage variant are limited. Of those, 63 originated from Ontario, 1 each originated from Quebec and British Columbia, and 1 originated from an unknown province or territory. In Canada, 66 R.1 lineage sequences were recorded, according to GISAID ( ), during December 2020–November 2021 ( 4). Most infections with R.1 variants have been reported in Japan and the United States ( 5, 6). In April 2021, the World Health Organization positioned R.1 lineage variants under the variant under monitoring (VuM) category to prioritize monitoring the variants because of distinct mutations in their genome. 
Globally, the R.1 lineage began to increase in frequency in December 2020, peaked in April 2021, became rare by June 2021, and was last reported in December 2021 ( 4). In Canada, the circulation of R.1 lineage variants corresponded with the third wave of the pandemic and was preceded by previously circulating variants of concern (VoC), B.1.1.7 (Alpha) and B.1.351 (Beta). We isolated SARS-CoV-2 R.1 lineage variants from an outbreak in persons facing housing insecurity in Hamilton, Ontario, Canada. Thus, continuously isolating and characterizing emerging SARS-CoV-2 variants is critical for developing updated vaccines and drug regimens. Mutations in the SARS-CoV-2 genome and within the spike glycoprotein alter the transmission dynamics, severity of disease, and sensitivity to neutralizing antibodies for each new variant ( 3). Although the first 2 waves were dominated by ancestral viruses, each subsequent wave had a surge in escape variants ( 1, 2). Since the beginning of the COVID-19 pandemic, Canada has encountered 8 waves of infections. SARS-CoV-2 continues to evolve and generate new variants. This critical driving force will influence the trajectory of the pandemic. Our study demonstrates that the R.1 variant retained sensitivity to neutralizing antibodies but evolved resistance to type I interferons. Of note, the R.1 variant was significantly more resistant to type I interferons (IFN-α/β) than was the ancestral isolate. The R.1 isolates were potently neutralized by both the wave 1 and wave 3 convalescent serum samples, unlike the B.1.351 (Beta) variant of concern. We used convalescent serum samples from persons in Canada infected either with ancestral virus (wave 1) or the B.1.1.7 (Alpha) variant of concern (wave 3) for testing neutralization sensitivity. In this study, we isolated samples of the SARS-CoV-2 R.1 lineage, categorized as a variant under monitoring by the World Health Organization, and evaluated their sensitivity to neutralizing antibodies and type I interferons. Isolating and characterizing emerging SARS-CoV-2 variants is key to understanding virus pathogenesis.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
